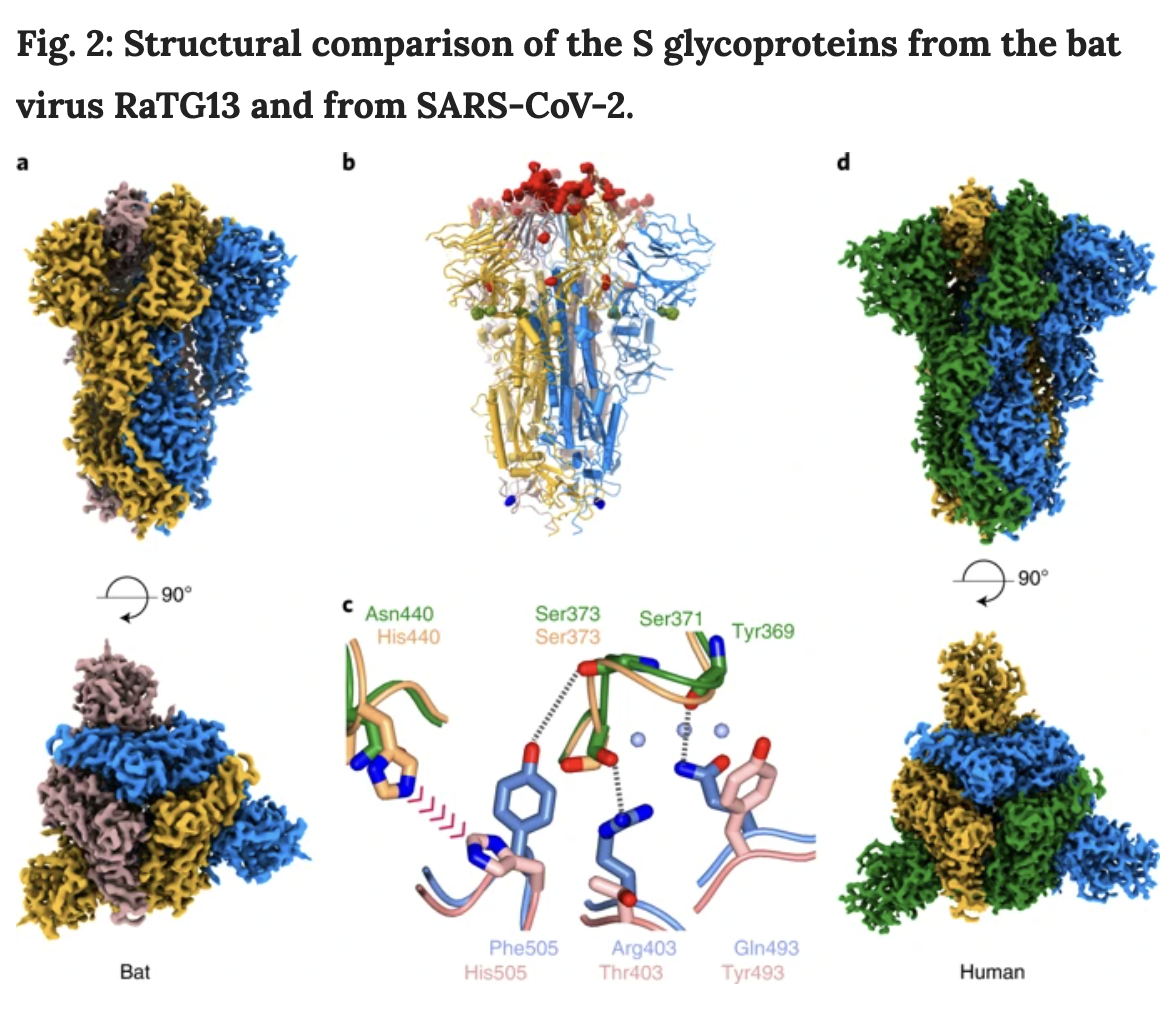
Fig. 2: Structural comparison of the S glycoproteins from the bat virus RaTG13 and from SARS-CoV-2
"a, The density map for the bat virus trimeric S is shown, with the long axis vertical in the top panel and in an orthogonal view in the bottom panel. All of the particles are in the closed conformation, likely because of the cross-linking of the material. The three monomers are colored blue, yellow and brown. b, Molecular model of the bat virus S protein, colored as in a, with substitutions between the bat virus and SARS-CoV-2 highlighted. Most of the changes are in the RBD and are colored red; there are four substitutions in S1 outside of the RBD, which are shown in green, and a single substitution in S2 is shown in blue. c, Overlay of the molecular structure of a portion of the RBD–RBD interface; the two bat virus S monomers are colored gold (top) and pink (bottom), and the two superposed SARS-CoV-2 S RBD chains are shown in green (top) and blue (bottom). Analysis suggests that the residues at the interface of SARS-CoV-2 S RBD chains support several additional stabilizing interactions and avoid the potential steric repulsion between His505 and His440, seen in the structure of the bat virus. d, The density map for the uncleaved SARS-CoV-2 S protein, in the closed conformation, shown in the same orientation as in a, with the subunits colored blue, green and yellow. This sample gave the best quality maps and enabled the most extensive build of the polypeptide chain."
0
1
Tags
SARS-CoV-2 (COVID-19)
Biomedical Sciences