Nonsynonymous TMPRSS2 Variations by Population
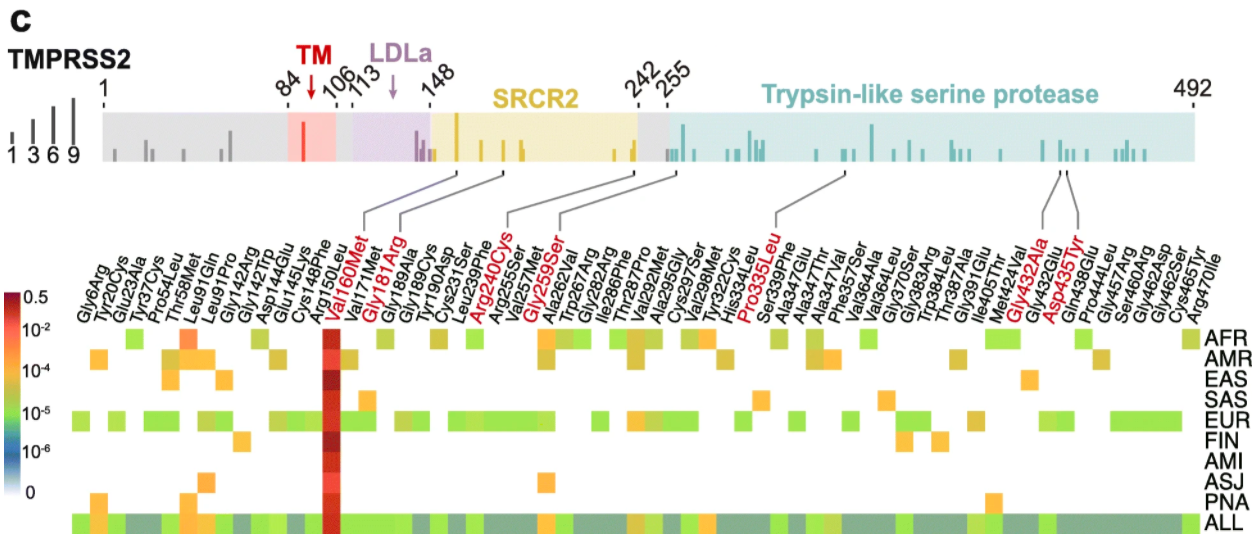
This graphic shows the distribution of 63 potentially harmful TMPRSS2 polymorphisms by population using variant assembly of databases, annotation of non-synonymous polymorphisms, and predictive-deleterious scoring.
- databases: Genome Aggregation Database, Exome Sequencing Project, and 1000 Genomes Project (1KGP, www.internationalgenome.org)
- non-synonymous variant annotation: ANNOVAR
- deleterious assessment: CADD and Polyphen2
The top bar shows TMPRSS2 functional domains, and the vertical notches represents population counts of the respective variant. The heat map depicts polymorphism frequencies found within the different populations.
- 35% of variations occur in African/African-American (AFR) populations
- 59% of variations occur in non-Finnish European (EUR) populations
- all populations carry p.Val160Met (~25% allele frequency)
- p.Asp435Tyr, which codes for a key residue for the substrate binding of TMPRSS2, is only carried by non-Finnish European (EUR) populations
- if TMPRSS2 polymorphisms cause gene overexpression, Down Syndrome may be a risk-factor for COVID-19 due to TMPRSS2's localization on chromosome 21.
Population group legend: AFR, African/African-American; AMI, Amish; AMR, Latino/Admixed American; ASJ, Ashkenazi Jewish; EAS, East Asian; FIN, Finnish; EUR, Non-Finnish European; SAS, South Asian; PNA, population not assigned
0
1
Contributors are:
Who are from:
Tags
SARS-CoV-2 (COVID-19)
Biomedical Sciences
Related
Age-Related Polymorphisms of TMPRSS2
Hypothesized Combination Therapies to Treat COVID-19 Based on Pharmacogenomics
Genetic variation within the TMPRSS2 protease may play a role in COVID-19 infection
Nonsynonymous ACE2 Variations by Population
Nonsynonymous TMPRSS2 Variations by Population
ACE2 variants and COVID-19